



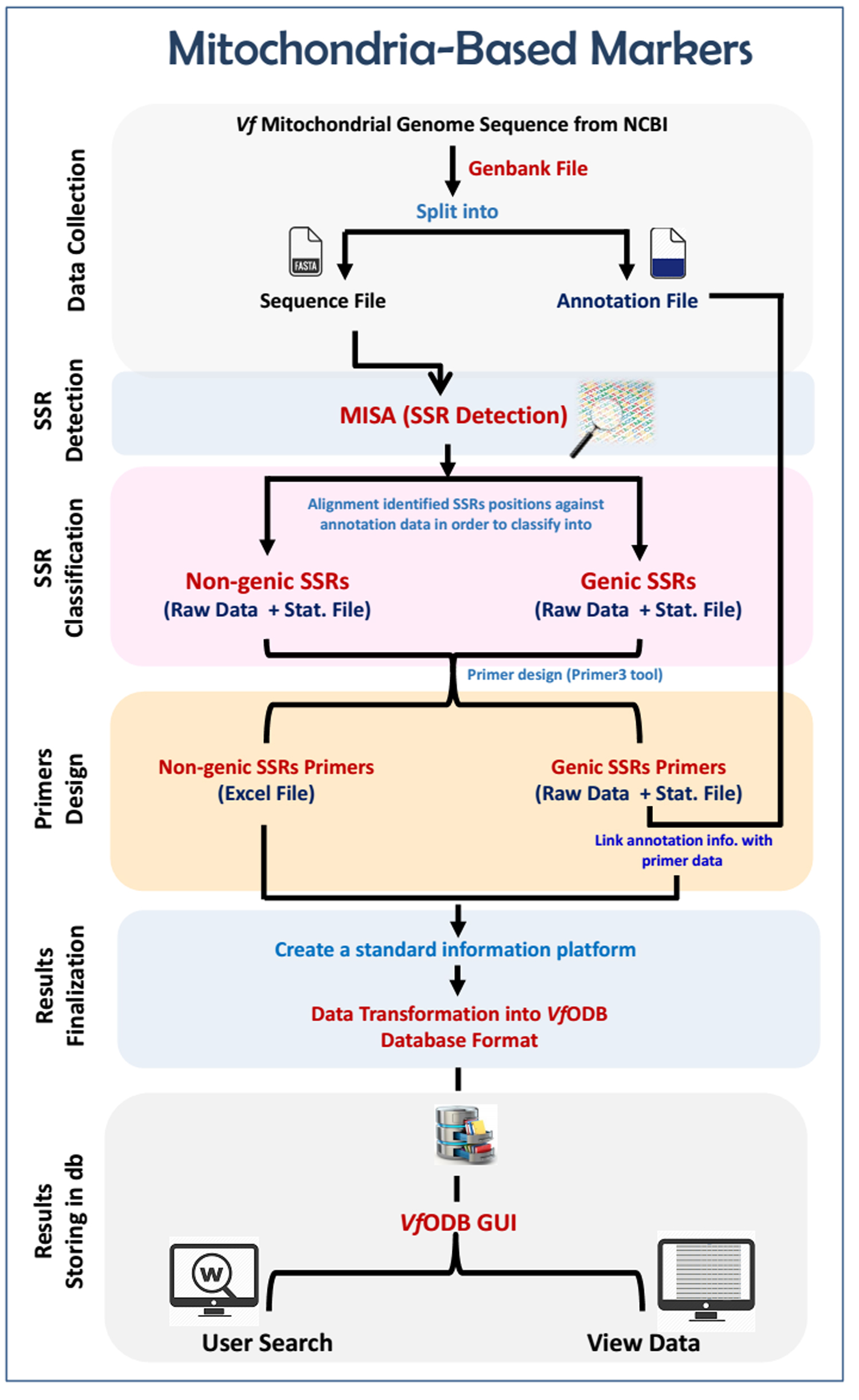
Vicia faba mitochondrial genome was analyzed in terms of microsatellites identification, classification and marker development. The workflow of analysis was done in five main steps, as follows: (a) detection of SSR motifs on the genome-scale; (b) classifying identified SSR motifs into non-genic or genic according to their location within the genome; (c) designing SSR primers and developing markers; (e) organizing and integrating developed SSR markers as well as their associated information into the VfODB database; and (f) implementing all generated datasets into the VfODB web interface
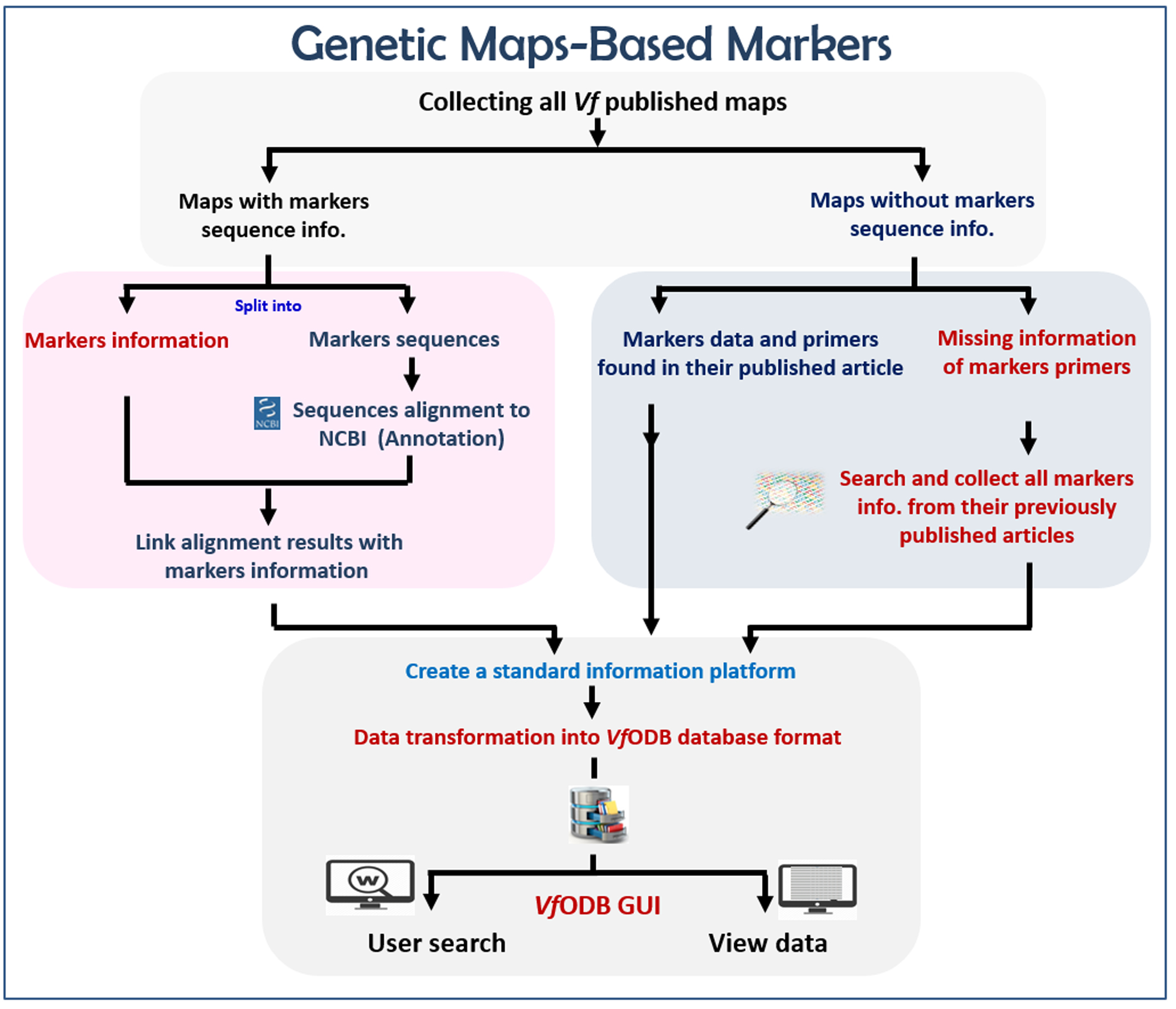
We downloaded all previously developed and published genetic linkage maps of Vicia faba. All available information of mapped markers on these maps were manually curated and categorized according to their type. In case that markers sequence was available such as SNP markers sequence, these sequences were further curated and aligned against the NCBI GenBank database by using blastx tool to determine its corresponding protein
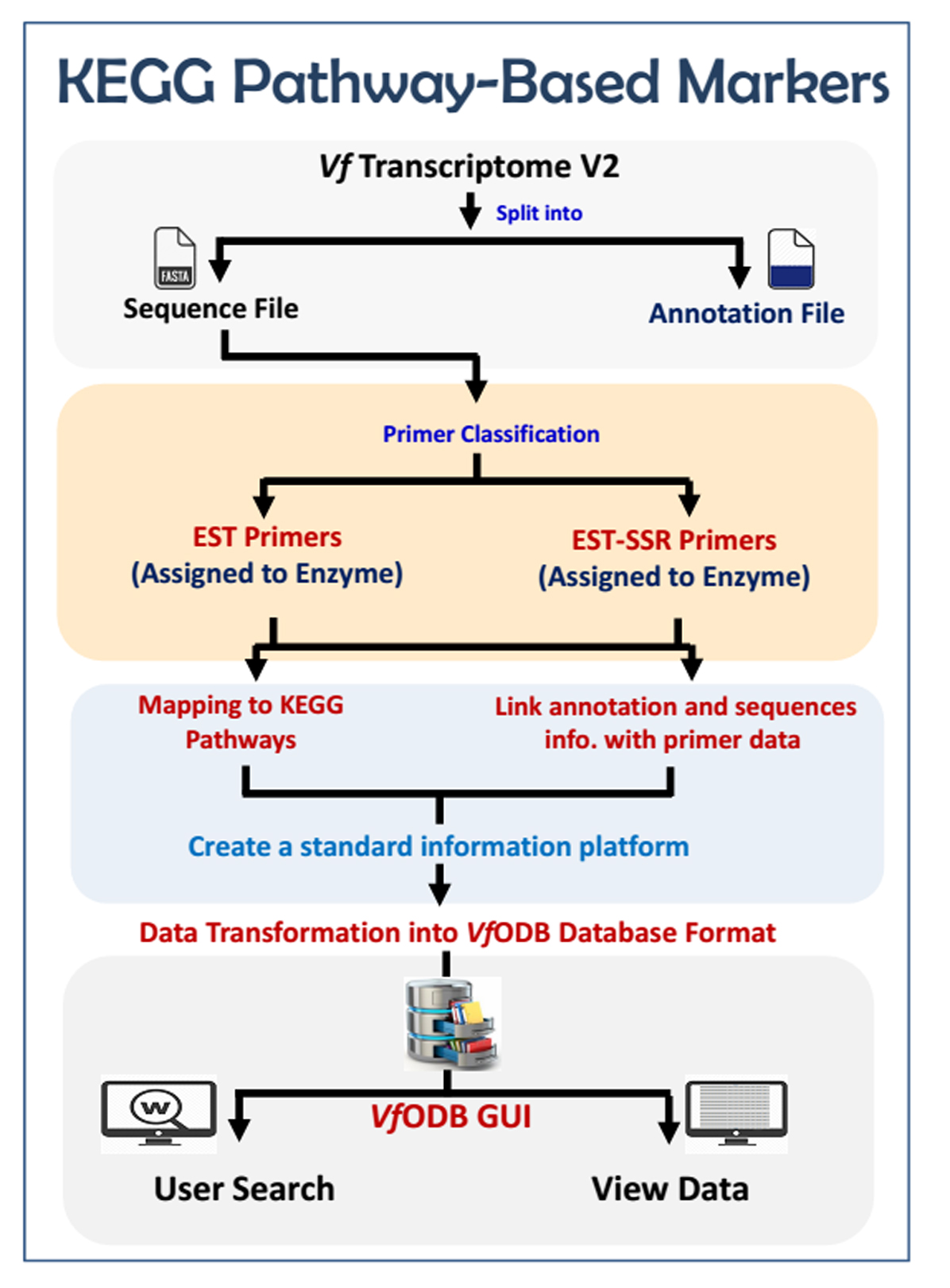
The developed EST and EST-SSR markers (assigned to enzymes) were further mapped against the KEGG pathways database to build Vicia faba functional maps. Each map contains; Pathway ID, Pathway image, mapped enzymes ID (highlighted), mapped enzymes associated primer/marker, marker annotation information and all other information related to this primer (Tm, GC%, Length, …etc.). All generated maps were finalized in a user-friendly attractive form.

Vicia faba mitochondria available in NCBI GenBank database (Accession number: KC189947)

Vicia fabatranscriptome version 2 was downloaded from Pulse Crop Database

Previously published Genetic Maps available in PubMed and Google Scholar till 2020

Our developed markers were further mapped against the KEGG pathways to build V. faba functional maps

We provide users with preliminary information about most of the faba bean germplasm available worldwide and listed in IPK Genebank and Genesys databases
